Why DNA Profiling?
The ability to extract and create a DNA profile is one of the most valuable tools in forensic investigation. It allows a sample of biological material (such as blood, semen, tissue) collected from a crime to be almost unequivocally linked to the person of whom it originated from.
Individuality
Due to polymorphic nature of DNA, the probability of finding two people with the same genetic profile is incredibly unlikely, therefore if a sample profile is matched to a profile of a suspect the chances are that they are most likely the origin of the sample. A staple principle in forensic science is that of Locard’s Exchange Principle, which states that every contact leaves a trace. This means that wherever someone goes they leave some type of trace of them having been there, which can materialise in the form of biological evidence left behind by the individual at a particular location. Thus, the discovery of an individual’s DNA at a scene can only place that individual as having been there within recent times but its presence alone is not sufficient to be considered proof of guilt to the crime committed.
The Beginnings of DNA Profiling
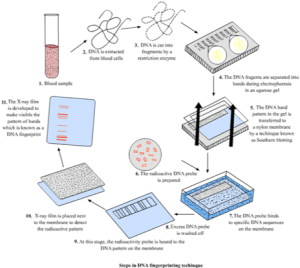
The first successful use of DNA profiling occurred in 1984 when English geneticist Dr. Alec Jeffreys used a technique known as restriction fragment length polymorphism (RFLP) to break DNA into smaller fragments for analysis. Jeffreys discovered that variable number tandem repeats (VNTRs) – which are structural regions of DNA where a short sequence of nucleotides is replicated a variable number of times in tandem – can be used as a genetic marker. What makes these repeat patterns within DNA so unique is that they vary in base length and number of repetitions for each individual person, therefore while many people may share the same pattern repeat, the way in which it is replicated will differ.
First successful Use
This new technique proved vital following the murder of two schoolgirls in Leicester, England. Semen samples were found and collected from both murder scenes, using Alec Jeffrey’s DNA profiling technique they were conclusively proved to be from the same individual. Two milestones were made following this discovery; a young boy who had originally been arrested for the murders after confessing to one but denying the other was the first person to be exonerated on the basis of DNA evidence, while the true guilty party, Colin Pitchfork, was the first man to be convicted of murder based on DNA evidence.
Since 1984 there has been great scientific advancements in the world of forensics, one of which is the more modern use of a short tandem repeat (STR) technology as a replacement for VNTRS in DNA testing. Both techniques share fundamentally similar principles however STRs work by analysing smaller lengths of DNA which makes them far more useful in cases where a DNA sample might be degraded – for example in cases of mass disasters where biological evidence has been exposed to extreme conditions. According to the Interpol Disaster Victim Identification Guide, DNA profiling is considered to be one of the primary methods in identifying missing and unknown persons following a disaster.
Unfortunately, most common, chemical forensic investigation methods destroy the DNA on exhibits of criminal acts. Therefore, EVISCAN provides a big advantage for the user in comparison to other methods for preserving traces. Here, a possible DNA will not be destroyed while the fingerprint is preserved and can also be secured. Therefore, a decision between preserving a fingerprint and a DNA trace is not necessary.
- Langford ,A., Dean, J., Reed, R., Holmes, D., Weyers, J., and Jones, A. (2005) Practical Skills in Forensic Science. 1st Pearson Education. UK.
- le.ac.uk. (2018) The history of genetic fingerprinting – University of Leicester. [online] Available at: https://www2.le.ac.uk/departments/genetics/jeffreys/history-gf
- Lynch, M. (2003). God’s signature: DNA profiling, the new gold standard in forensic science. Endeavour, 27(2), pp.93-97.



